Recently, our laboratory obtained a high-quality reference genome of tea plant (Camellia sinensis var. sinensis) at the chromosome level and revealed the mechanism of adaptive evolution of cultivated tea plants. The research results were published in the international botanical journal Molecular Plant, named "The reference genome of tea plant and resequencing of 81 diverse accessions provide insights into genome evolution and adaptation of tea plants"(IF=10.8). This is another major landmark research created by the research team of Wan Xiaochun and Wei Chaoling in the field of basic tea biology after the publication of genome draft of Chinese tea plant species (PNAS, 2018, 115(18): E4151-E4158) and the successful construction of a comprehensive genomics and bioinformatics platform for tea plant. (Plant Biotechnology Journal, 2019, 17(1): 1938-1953).
With the national tea tree variety Shuchazao (Chinese species) as the material, this study used single-molecule sequencing (PacBio) and chromosome conformation capture (Hi-C) technologies to overcome the assembling difficulties in large amount, high heterozygosity and repetition of tea genomes. As a result, it obtained high-quality reference genome sequence of tea trees at the chromosome level, which has Contig N50 of 600 Kb, Scaffold N50 of 167 Mb and BUSCO completeness of 94%. The accuracy and completeness of this genome assembly were greatly improved compared with the previously reported tea plant genome draft. On this basis, comparing genomics and population genetics studies revealed that:
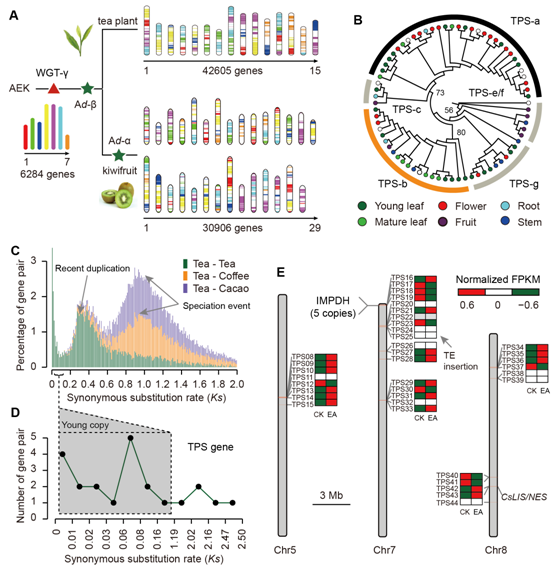
1)The high content of repetitive sequence is not only the main reason for the large size of the tea plant genome, but also allows the increase of the average gene length and the functional differentiation of some repetitive genes through intron insertion;
2) The heterozygous region of the tea tree genome accounts for 18.8% of the whole genome, and contains 3,440 protein-coding genes, which are mainly involved in biological processes such as nitrogen compound transport activity, histone modification and hormone synthesis;
3) Terpene synthase genes related to tea aroma and resistance were found to be significantly amplified mainly by recent tandem repeat events and distributed in the tea genome mainly in the form of gene clusters in different chromosomes;
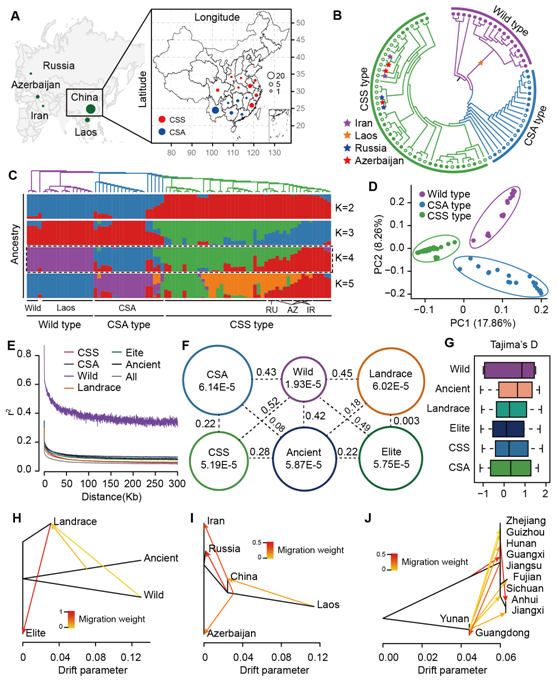
4) Through deep sequencing of 81 representative tea samples at home and abroad, the first genomic variation map of representative cultivated and wild-growing tea plants was framed, and it was found that the selected samples were clearly classified into Assam, Chinese species and wild type; The results of genetic diversity studies of tea plants from different regions of China supported the theory of southwestern origin of cultivated tea trees in China; Some artificially acclimated genes that were strongly selected during the selection and improvement of tea varieties were identified. The research results will provide high-quality data resources and theoretical basis for the future scientific conservation of excellent germplasm resources of tea trees, gene discovery of important agronomic traits of tea trees, development and utilization of health and efficacy components of tea leaves and genetic breeding research, and will also further promote the research process of major basic biological issues such as genome evolution of Camellia sinensis, origin and genetic diversity of tea trees, and formation mechanism of characteristic secondary metabolites of tea leaves. It will also promote the understanding, dissemination and utilization of tea in the world.
Copyright 2015 State Key Laboratory of Tea Plant Biology and Utilization All Rights Reserved